Simon Goring – University of Wisconsin – Madison
Problem Statement
Complexity is a significant hurdle to user adoption of new tools and workflows (Kim and Crowston, 2011). Complexity also leads to heterogeneity in data processing and archiving practices (workflows), as best practices are often culturally transmitted. In the “long tail” of science, data (which includes scientific workflows) tend to be collected and generated for project-specific purposes, with greater emphasis on sharing within research groups than with the public (Wallis et al., 2013). Heterogeneity can then limit our ability to undertake synthesis work, and complicates the transmission of knowledge, particularly when disciplinary communities are small and highly distributed, as in the paleogeosciences. Open publication of results and data may be insufficient for limiting these challenges given the complexity of many modern workflows, thus a disparity will continue to exist between the standards for publication, with extensive peer review, and the standards for data archiving (Lee and Bietz, 2009).
To solve the challenge of best-practices heterogeneity around new tools & techniques, I propose a system, rNoteBase, to archive and expose workflows generated using Rmarkdown (focusing on API-based tools supported by rOpenSci). Documents linked to rNoteBase can be annotated, tagged with keywords, and edited through existing version control platforms (e.g. GitHub). rNoteBase will not host the documents directly, rather linking to the documents from a code repository, this helps balance centralizing tendencies of technology with the distributed nature of modern scientific collaboration (Lynch, 2008).
The rNoteBase will assist researchers move toward greater transparency by facilitating reproducibility and increasing the competency of researchers for core technical skills. One barrier for code sharing is the sense that code or data is too messy (Barnes, 2010). By providing researchers with tools, written with scientific and technical best practices in the forefront, we encourage reproducibility. By basing initial generation of this system around the API-focused rOpenSci packages we hope to foster partnerships between data creators, data repositories, and data users.
This project was originally a component of the NSF funded Throughout proposal but was removed due to funding constraints. It is my belief that the rOpenSci fellowship would provide the opportunity to build a working prototype of this platform in connection with work underway in the Throughput proposal that is designed to link resources across the geoscientific web, as part of the EarthCube program (http://earthcube.org).
Proposed Activities and Outcomes
rNoteBase will ensure (1) that workflows undergo a vetting process; (2) that end-users are assured that web-centric methods are reliable; (3) that workflows contained within the notebase are discoverable, and (4) that credit can be provided to rNoteBase contributors. To accomplish this, the rNoteBase proposal consists of four elements:
A Peer Review System for API-centered scientific workflows that leverages the existing rOpenSci peer-review system (e.g., https://github.com/ropensci/onboarding), but remains separate and domain-specific to avoid overburdening the existing system.
A Continuous Integration System to check notebooks regularly (cf., CI with Rmarkdown), to provide failure notification to notebook developers and, if requested, API maintainers.
A User Interface, leveraging RMarkdown YAML headers to help sort and manage notebooks, providing search capacities and allowing discovery by keyword, discipline, R package, or author. The intention is to construct a system using an API-first philosophy, with vue.js to support the UI development. This rNoteBase would be able to support its own rOpenSci package.
A DOI Management System to be supported initially through the University of Wisconsin’s Library System and its contract with DataCite along with data citations for the workflow products to support recognition.
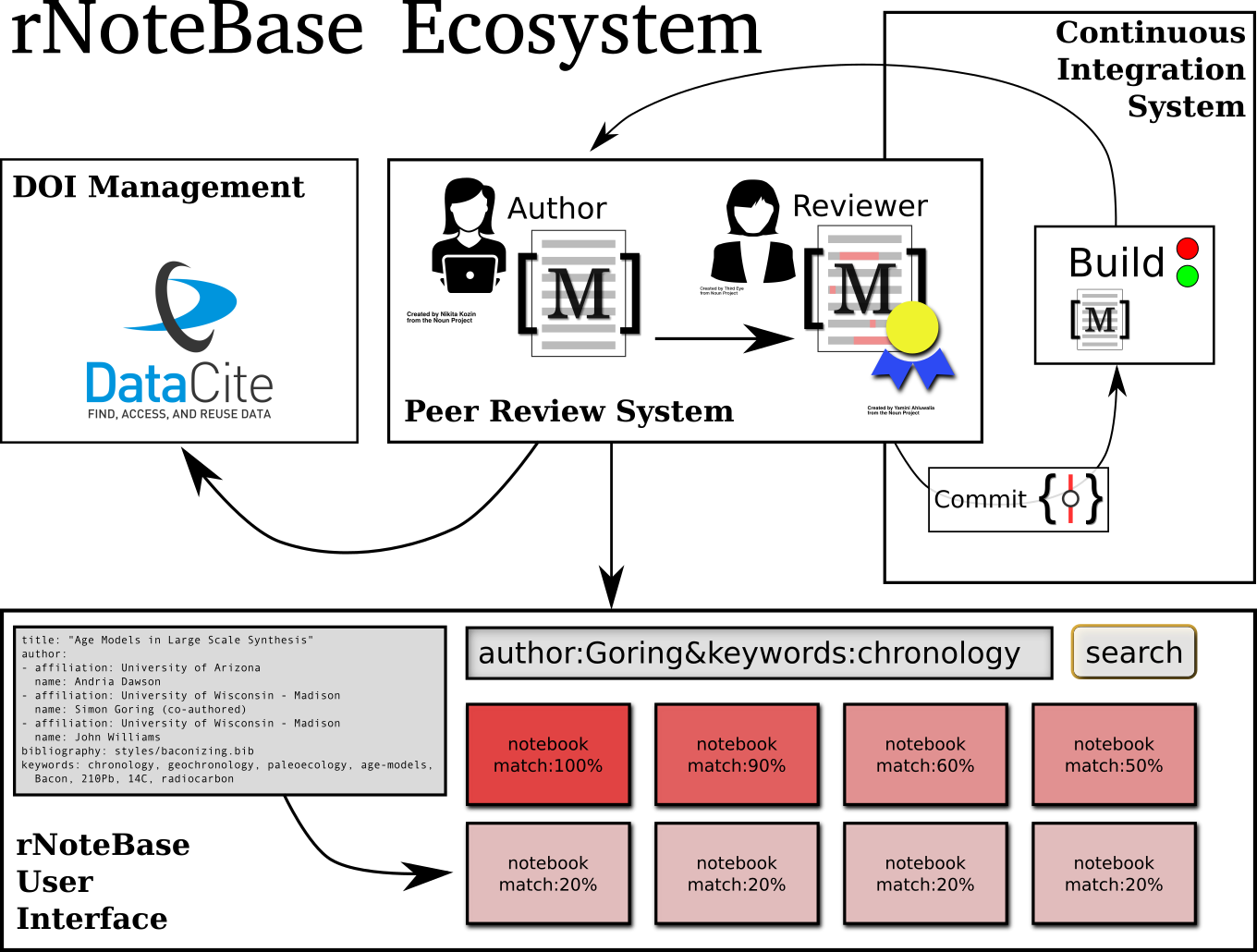
Figure 1: Visual representation of the elements within the rNoteBase ecosystem. Individual elements are described further in the proposal.
The implementation plan for rNoteBase differs from the more common development of end-user tools in that it will provide descriptive, citable workflow resources for researchers engaged in domain research. This ensures that notebook authors recieve academic credit through citations, and that users receive support by having peer-reviewed general workflows to use as a type of “cookbook”.
Populating the Resource
Workflows developed with the Neotoma Paleoecological Database (http://neotomadb.org) include questions critical to the paleogeosciences broadly, such as chronology construction using Bayesian tools (e.g., based on http://bit.ly/2mkAVCd), age-uncertain analysis of paleogeoscientific data (using GeoChronR tools; McKay), sediment core alignment using affine and splice, geospatial-temporal mapping of lake water chemistry, or sediment geochemistry from EarthChem, pollen-based climate reconstruction (e.g., http://bit.ly/2m1clUx), vegetation reconstruction using pollen, cluster analysis of tephra events, or sample mapping and correlation of paleo-proxies. We will use these as the initial basis for document links, metadata harvesting and annotation. As the platform approaches maturity we wil solicit additional workflows through social media channels and relationships with partners in the EarthCube program as part of the EarthCube Engagement Committee’s work. We will also work to harvest information from existing package vignettes that perform discipline-related tasks.
Tentative Timeline
This proposal would result in several deliverable products. All code for the prototype would be available on GitHub under an MIT License. Content would be dynamically linked, and as such, the rNoteBase would not host markdown documents itself.
First Three Months
Standards for data objects defined from DublinCore, WC3, schema.org and other ontologies
Data model for DOI atomic units & data citation elements
First Six Months
Prototype neo4j graph database linking RMarkdown documents to user profiles (e.g., ORCiD), peer reviewers, API resources and R package information
API developed using vue.js to query the graph database and return content based on rNoteBase workflow attributes (e.g., keywords, packages used, data types accessed, authors)
First Nine Months
Peer review process outlined, discussions with potential partner organizations (Ubiquity Press, Earth Science Information Partners – currently engaged in development of an earth science preprint service)
First Year
A web application developed using vue.js
Biography
Simon Goring (http://goring.org) is an Assistant Scientist at the University of Wisconsin-Madison, with PI status. He is the lead author of the neotoma R package and the IT lead for the Neotoma Paleoecological Database, as well as a member of the Leadership Council for the NSF’s EarthCube Program.
We aim to leverage resources from the NSF Throughput grant to help undertake a workshop at the University of Wisconsin in the summer of 2018 that would focus on workflow development for the paleogeosciences. This would provide further support to the rNoteBase proposal by adding content and acting as an opportunity to beta-test the platform.
References
Barnes, N. (2010). Publish your computer code: it is good enough. Nature, 467: 753. doi:10.1038/467753a
Kim, Y., & Crowston, K. (2011). Technology adoption and use theory review for studying scientists' continued use of cyber‐infrastructure. Proceedings of the Association for Information Science and Technology, 48:1-10. doi:10.1002/meet.2011.14504801197
Lee, C. P., & Bietz, M. (2009). Barriers to the adoption of new collaboration technologies for scientists. In: ACM Conference on Computer-Human Interaction (CHI). [link]
Lynch, C. (2008). The institutional challenges of cyberinfrastructure and e-research. Educause Review, 43: 74-88. [link]
Wallis, J. C., Rolando, E., & Borgman, C. L. (2013). If we share data, will anyone use them? Data sharing and reuse in the long tail of science and technology. PloS one, 8(7), e67332. doi:10.1371/journal.pone.0067332